Research Overview
Our research focuses on pathological and functional aspects of protein disorder and aggregation. In particular, we are interested in the differences between functional and pathological cross-β (amyloid) fibrils. Further, we want to determine the role of intrinsically disordered protein domains (i.e. domains that lack stable tertiary or secondary structure) that are often found in these fibrils for both function and toxicity.
Cross-β fibril formation is important for many neurodegenerative diseases such as Alzheimer's, Parkinson's, and Huntington's disease. The aggregation of monomeric proteins into a variety of oligomeric and fibrillar (amyloid) states can lead to cell death and, therefore, disease. However, cross-β fibrils can also have non-pathological, positive functions such as cell signaling or scaffolding. One of our goals is to determine what distinguishes functional and pathological cross-β fibrils. Knowing the difference between these types of fibrils can potentially lead to the development of new therapies and will deepen our understanding of the role of cross-β fibrils in biology.
 Electron microscopy image of fibrils formed by the functional amyloid Orb2 important for long-term memory
Electron microscopy image of fibrils formed by the functional amyloid Orb2 important for long-term memory
Another focus of our research is intrinsic protein disorder that is often found in cross-β fibrils. In recent years it has become clear that intrinsically disordered proteins carry important functional aspects, especially in eukaryotic organisms, where up to 40% of all proteins contain disordered regions. Recent work by our team at USC and others has shown that the intrinsically disordered domains of huntingtin exon-1 fibrils important in Huntington's Disease dominate the surface of these fibrils and might be key for understanding fibril toxicity. Therefore, our goal is to determine the conformational ensemble of these disordered domains and their specific interaction with other cellular components.
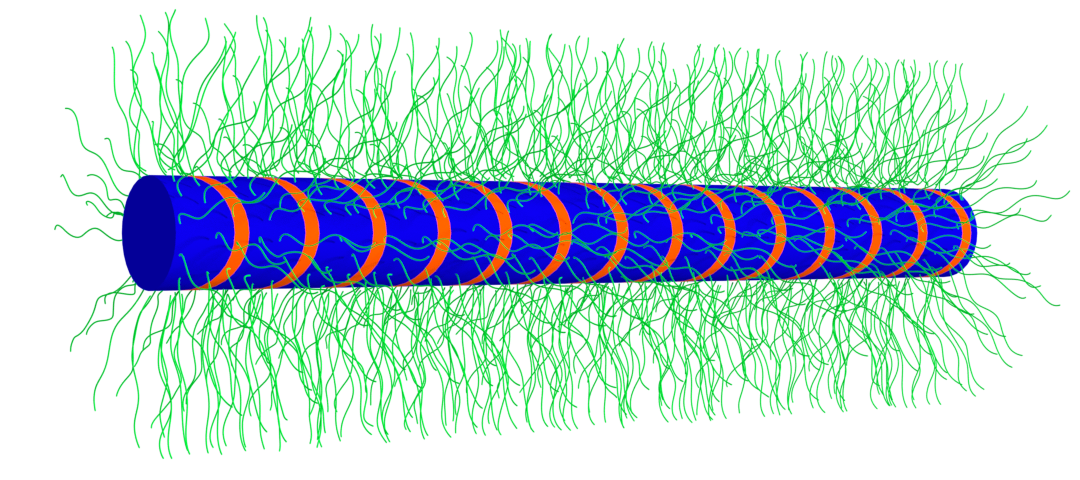 Bottlebrush model of cross-β fibrils with central cross-β core and dynamic framing sequences at the surface
Bottlebrush model of cross-β fibrils with central cross-β core and dynamic framing sequences at the surface
To address these questions, my lab will use primarily solid-state nuclear magnetic resonance (NMR) in conjunction with other biophysical methods. Solid-state NMR enables us to investigate protein structure and especially protein dynamics in non soluble, non-crystalline, non-frozen states, which allows us to study the structure and dynamics of these aggregates with atomic resolution.